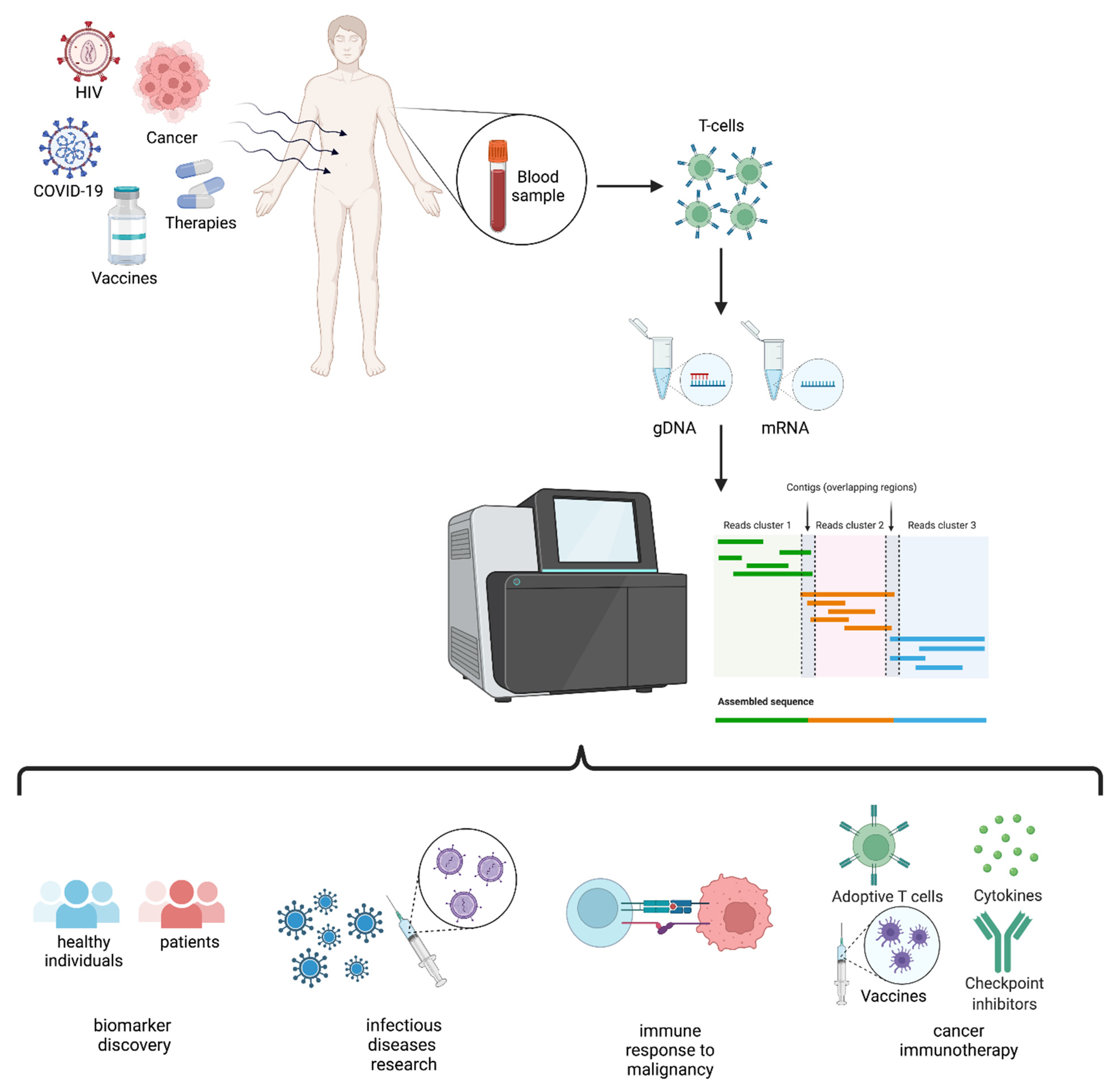
There are several approaches for extracting CDR data from sequencing reads and determining the clonotype. 7, 18 This approach has provided an estimation of approximately 10 6 unique TCR sequences present in a human individual. 16, 17 Recent next-generation sequencing (NGS) technologies have offered significant advantages over these earlier methods and have enabled extensive, comprehensive high-throughput measurement and analyses of the diversity of both TCRs and B-cell receptors (BCRs) to elucidate immune functions in health and disease. Using techniques such as Southern blot hybridization, polymerase chain reaction and flow cytometry, researchers have characterized T-cell proliferation. In addition, quantitative assessment of T-cell clones can be of important prognostic significance to manage patients and their treatment. 8 Analysis of the TCR repertoire can thus characterize the host T-cell immune status in patients and normal individuals. 12, 13, 14 In contrast, clonal populations are a hallmark of malignancy, and clonal or oligoclonal populations of T and B cells can also occur in non-malignant conditions including human immunodeficiency virus and Epstein–Barr virus infections. 11 In normal individuals, the TCR repertoire is polyclonal and non-static. 10 The clonotype of a TCR population is a molecular description of the unique sequences (typically described from the amino acid perspective) required to produce the antigen specificity of TCRs, as well as the specific V and J genes involved in the composite rearrangements. The ability of human T cells to recognize a vast range of pathogens and initiate specific adaptive immune responses depends on the versatility of the TCR population. 1, 9 Thus, the sequence of CDR3 and the identity of the flanking V and J gene segments are widely used to classify TCR variants. 7, 8 The vast majority of TCR variation is within complementarity-determining region 3 (CDR3), which encompasses the VDJ recombination junctions and encodes the portion of the TCR that directly contacts peptide-bound major histocompatibility complex molecules. 3, 6 Because this somatic rearrangement occurs only in the T-cell genome and produces an extremely diverse repertoire of TCRs as a hallmark of the cellular adaptive immune system, it can be effectively used as a unique tag to enumerate and quantify T-cell clonality. 4, 5 An individual’s TCR repertoire is shaped by biases during the process of VDJ recombination and by the subsequent expansion and deletion of certain T-cell clones upon antigen recognition during T-cell development in the thymus and later in the periphery. 3 The TCR consists of a heterodimer of two chains (αβ or γδ), both of which are products of V(D)J recombination. 1, 2 This process is directly analogous to the generation of antibody diversity by somatic VDJ recombination of the B-cell receptor locus in B lymphocytes. The T-cell receptor (TCR), which is generated through random rearrangement of genomic V(D)J-variable(diversity)joining-segments, is the mediator of specific antigen recognition by T lymphocytes. Thus, determining monoclonal architecture and clonal diversity by RNA sequencing might be useful for prognostic purposes and for personalizing ATL diagnosis and assessment of treatments. CDR3 sequence length distribution, amino acid conservation and gene usage variability for ATL patients resembled those of peripheral blood cells from ACs and healthy donors. ACs, CD4 -positive samples and smoldering patients showed significantly higher TCR diversity compared with chronic, acute and lymphoma subtypes. Expression of TCRα and TCRβ genes in the dominant clone differed among the samples. Based on these TCR profiles, CD4-positive cells and ACs showed polyclonal patterns, whereas ATL patients showed oligo- or monoclonal patterns (with 446 average clonotypes across samples). We analyzed 62 sets of high-throughput RNA sequencing data from 59 samples of HTLV-1−infected individuals-asymptomatic carriers (ACs), smoldering, chronic, acute and lymphoma ATL subtypes-and three uninfected controls to evaluate TCR distribution. Here we examined the diversity of TCR clonality and its association with pathogenesis and prognosis in adult T-cell leukemia/lymphoma (ATL), a malignancy caused by infection with human T-cell leukemia virus type-1 (HTLV-1). The TCR repertoire can be altered in the context of infections, malignancies or immunological disorders. High-throughput sequencing technologies now allow examination of antigen receptor repertoires at single-nucleotide and, more recently, single-cell resolution. The diversity of T-cell receptor (TCR) repertoires, as generated by somatic DNA rearrangements, is central to immune system function.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
